Assistant Professor Dinler Amaral Antunes Publishes Study Outlining Tool Development
In an effort to develop drugs to treat COVID-19, a team of multidisciplinary researchers created an online tool that can predict whether a drug molecule can be docked to a protein on the SARS-CoV-2 virus.
Dinler Amaral Antunes, assistant professor of computational biology at the University of Houston, along with scientists from Rice University, Scotland’s University of Edinburgh and Brazil’s Federal University of Ceará, collaborated to create the webserver DINC-COVID. They published their work in the journal Computers in Biology and Medicine.
DINC, which stands for Docking INCrementally, is an online tool that can screen whether a drug can bind to a protein that’s being targeted for treatment. Antunes helped develop the tool while he was a postdoctoral researcher in the lab of Rice University computer science professor Lydia Kavraki.
At the beginning of the pandemic, Geancarlo Zanatta, a former collaborator from Brazil, who is now associate professor at the Federal University of Ceará, reached out to Antunes saying many researchers were attempting to run virtual screenings for drugs that could be used to treat COVID-19, without much success. Zanatta said one of the issues researchers were not considering was the flexibility of the protein receptor.
“X-Ray crystallography, the experimental method for protein structure determination, provides a ‘static picture’ of the protein structure,” said Antunes, faculty at UH’s College of Natural Sciences and Mathematics. “However, proteins have dynamic behavior in solution. They can oscillate between states and that has a direct impact on the shape of the binding sites available for drug interaction.”
For example, the team writes that although SARS-CoV-2’s Main protease (Mpro) has so far been one of the most explored coronavirus targets in computational studies, there are still many questions about the design of inhibitors or drugs to treat the virus. “The malleability of the Mpro active-site cavity remains the greatest challenge in the development of effective inhibitors,” the study’s authors write.
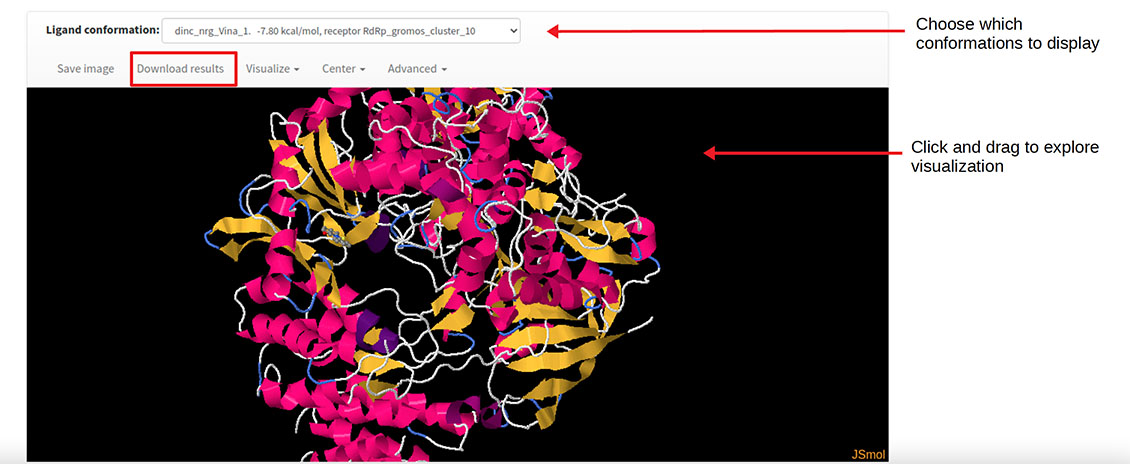
To account for the protein flexibility, the team made DINC-COVID, a version of DINC built specifically to test drugs’ ability to dock to multiple conformations of SARS-CoV-2 proteins. The webserver uses molecular dynamics simulations to produce a “movie” of the motions of proteins. Researchers then extract representative “pictures” of what was captured by the movie, and they offer this ensemble for users to test the binding of their drug candidates. Results are emailed to the users.
DINC-COVID Available to Users Without Computational Expertise
“More than 500 people have used the server from 54 different countries,” Antunes said. “There are people all over the globe using the tool right now for predictions and trying to find other protein binders.”
Simulating molecular dynamics is time consuming and requires high-performance computers. It limits the ability to dock multiple protein conformations to researchers who have these resources. Antunes is proud of the accessibility and ease their tool provides to people anywhere in the world with an internet connection. DINC-COVID enables both people with limited computational resources and people who do not have a computational background to test their own molecules, he said.
The paper’s authors note that although vaccines have been validated and used in massive vaccination campaigns, particularly in developed countries, “the need for pharmacological treatments for infected patients persists due to unequal vaccination coverage across the globe” and because of “the rise of more virulent variants that can cause symptoms even in fully-vaccinated individuals.”
Their challenge during the beginning of the pandemic was to hopefully create a solution with their expertise.
“Most of my work is basic research,” Antunes said. “I’m developing methods and potentially drugs that could come to market in several years, in the best-case scenario. Even with the vaccine results, there’s still room for drugs to help treat patients. That’s exciting. This is one case in which we will get close to make a direct impact, although we are on the prediction side.”
Other research groups can make their own predictions using DINC-COVID as a baseline and then test results in the lab to see if the findings are working as expected. Antunes is not aware of any drugs that have been developed using the tool yet.
Speedy Development Thanks to Broad Range of Subject Matter
The study included researchers from a variety of disciplines, ranging from biophysics to computer science to biochemistry.
“This enabled us to be slightly faster,” he said. “Many of these research topics are complex and too difficult for one person to tackle all aspects of it. It’s important to do these collaborations with people who have different expertise to make progress more quickly.”
Future plans for DINC-COVID include expansion of the number of proteins available to work with on the webserver.
The study was funded by the National Science Foundation, Brazil’s National Council for Scientific and Technological Development, and Rice University.
- Rebeca Trejo, College of Natural Sciences and Mathematics