Faculty Profile
 Erin Kelleher
Erin Kelleher
Associate Professor and Associate Chair for Graduate Affairs
Department of Biology and Biochemistry
Research Division: Cell and Molecular Biology (Primary), Ecology and Evolution (Joint/Adjunct)
Office: Science & Research 2, 421D
Contact: eskelleher@uh.edu - (713) 743-3640
Education: Ph.D., University of Arizona, B.S./B.A., University of Virginia
Google Scholar Profile
Website
The Kelleher lab is interested in the defense of the genome to sources of mutation including transposable elements, UV and ionizing radiation. We the molecular mechanisms that underlie genome defense, natural variation and evolution of these mechanisms.
UV and Ionizing Radiation
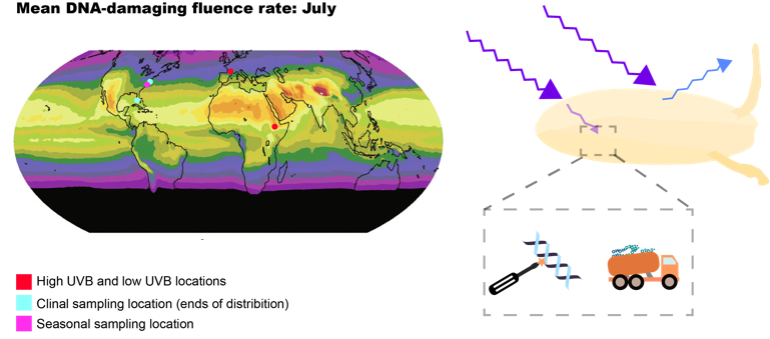
UV and ionizing radiation pose a major threat to genome integrity by producing mutations. Organisms defend their genomes against the harmful effects of radiation through physical barriers (UV only) and DNA repair. We are interested in how these defense mechanisms vary between individuals, and how this relates to geographic and seasonal variation in UV and ionizing radiation. This work is also relevant to radiation therapy, which relies on differential radiotolerance between healthy and tumor cells.
Transposable elements
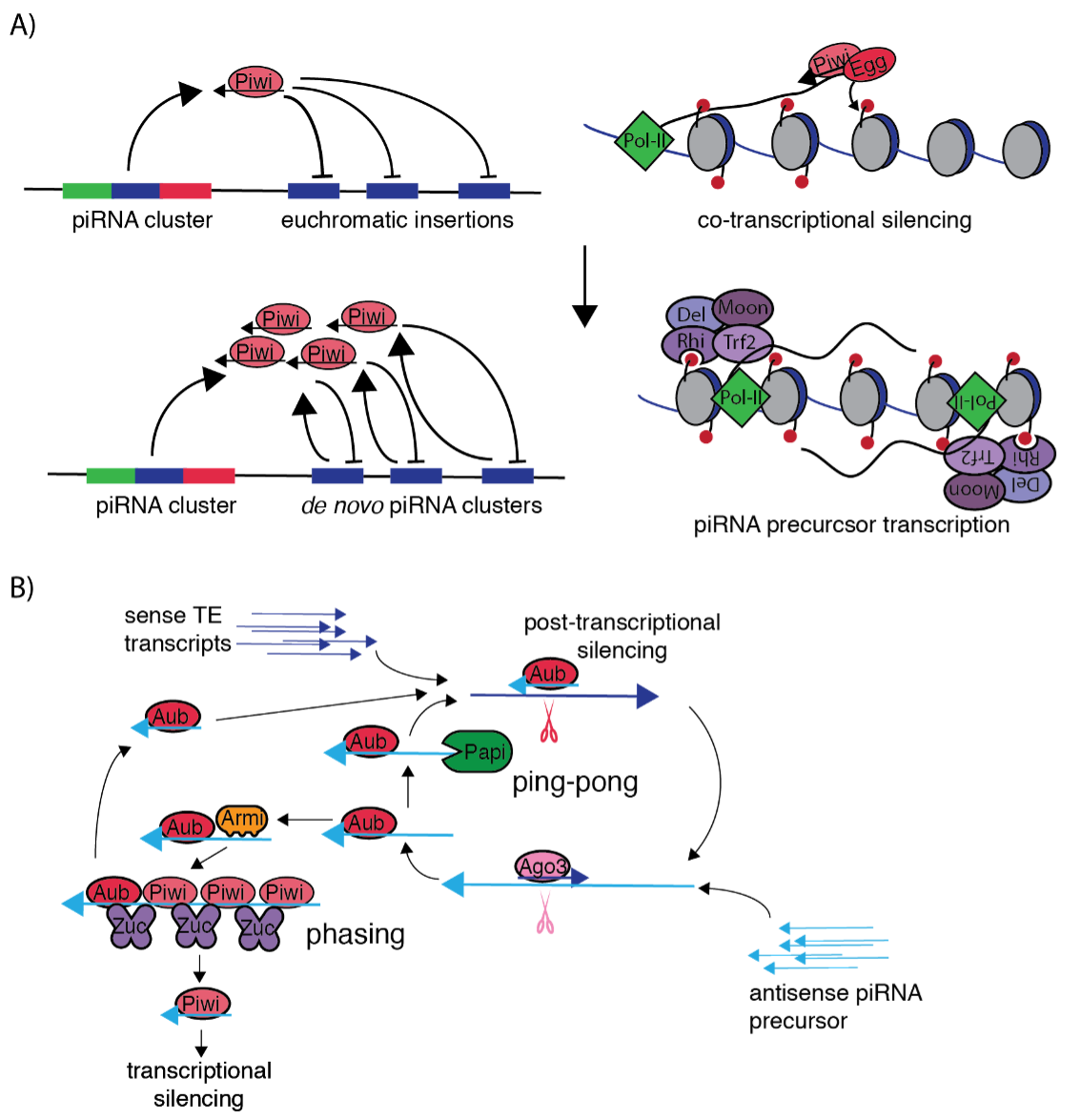
Transposable elements (TEs) are mobile genetic sequences that populate and parasitize genomes. TEs impose a multifaceted mutational burden on their host through insertional inactivation of host genes, participation in ectopic recombination, and the induction of DNA damage. Our research focuses on how the host genome fights back against TEs through small RNA silencing pathways, and how host facilitate the movement of transposable elements through transpositional cofactors.
- Kelleher E.S. Jack of all trades versus master of one: how generalist versus specialist strategies of transposable elements relates to their horizontal transfer between lineages. Curr Opin Genet Dev. 2023 81:102080. doi: 10.1016/j.gde.2023.102080.
- Wang L., Zhang S., Hadjipanteli S., Saiz L., Nguyen L., Silva E., and Kelleher E.S. 2023) P-element invasions fuels molecular adaptation in laboratory populations of Drosophila melanogaster. Evolution, 77:980-994.
- Lama, J., Hadjipanteli, S., Srivastav, S. , Tasnim, S., Hubbard, D.A., Smith, B.R, Maconald, S.J., Green, L. and Kelleher E.S. (2022) Genetic variation in P-element dysgenic sterility is associated with double-strand break repair and alternative splicing of TE transcripts. PloS Genetics, 18:e1010080.
- Kelleher E.S. (2021) Protein-Protein Interactions Shape Genomic Autoimmunity in the Adaptively-Evolving Rhino-Deadlock-Cutoff (RDC) Complex. Genome Biology and Evolution, 13:evab132.
- Wang L., Barbash D.A., Kelleher E.S. (2020) Adaptive evolution among cytoplasmic piRNA proteins leads to decreased genomic auto-immunity. PLoS Genetics, 16: e1008861.
- Kelleher E.S., Barbash D.A., and Blumenstiel J.P. (2020). Taming the Turmoil Within: New Insights on the Containment of Transposable Elements. Trends in Genetics, 36:474-489.
- Zhang S., Pointer B., Kelleher E.S. (2020) Rapid evolution of piRNA-mediated silencing of an invading transposable element was driven by abundant de novo mutations. Genome Research, 30:566-575.
- Kelleher E.S., Jaweria J., Akoma U., Ortega L., Tang W. (2018) QTL mapping of natural variation reveals that the developmental regulator bruno reduces tolerance to P-element transposition in the Drosophila female germline. PLoS Biology. 16:e2006040.
- Tasnim S., Kelleher E.S. (2018) p53 is required for female germline stem cell maintenance in P -element hybrid dysgenesis. Developmental Biology. 434:215-220.
- Kelleher, E.S., Barbash D.A. (2013) Analysis of piRNA-mediated silencing of active TEs in Drosophila melanogaster suggests limits on the evolution of host genome defense. Molecular Biology and Evolution. 30:1816-1829.
- Kelleher, E.S., Edelman, N.B., Barbash, D.A. (2012) Drosophila interspecific hybrids phenocopy piRNA pathway mutants. PLoS Biology. 10:e1001428.
Honors and Awards
National Institute of Health National Research Service Award, 2010-2013
Cornell Center for Population Genomics Post-Doctoral Fellow, 2009-2010
American Association for University Women Dissertation Fellowship, 2008-2009
National Science Foundation Doctoral Dissertation Improvement Grant, 2007
National Science Foundation IGERT Fellow in Genomics, 2004-2008
Mentorship Training
|
Understanding Neurodiverse Motivation and Learning Needs
|
2024 |
|
Culturally Aware Mentorship Facilitator
|
2023 |
|
Culturally Aware Mentorship
|
2021 |
|
Gulf Coast Consortium Mentor Training
|
2020 |
| Scientific Communication Enhances Research Excellence |
2020 |
Professional Service
|
Editor PLoS Biology
|
2022-present
|
|
NIH Panelist
|
2021, 2024
|
|
NSF Panelist
|
2015 |
|
Co-Chair – Evolution and Quantitative Genetics Session, Drosophila Research Conference
|
2015, 2019 |
|
Organizer – Southeast Texas Evolutionary Genetics and Genomics Symposium
|
2016, 2022, 2023 |
College Service
|
Graduate Affairs Committee
|
2021-present
|
|
Strategic Planning for Graduate Student Success
|
Spring 2022
|
|
Strategic Hiring Committee
|
Spring 2024
|
Outreach
| Research Experience for Teachers (RET) Sponsor (NSF) | 2016, 2017, 2019, 2021-present |
| Developer an Co-intstructor of the Drosophila Genetics Workshop for 7th Grade Science Teachers at the University of Houston | 2015, 2016, 2017, 2019 |